Category:Biased molecular dynamics
Biased molecular dynamics (MD) refers to advanced MD-simulation methods that introduce a bias potential.
For instance, umbrella sampling[1] uses a bias potential to enhance the sampling of a geometric parameter in regions with low probability density, e.g., transition regions of chemical reactions. The correct distributions are recovered afterward using an exact relation between the biased and unbiased systems. D. Frenkel and B. Smit provide a more detailed description of the method in Ref. [2]. Biased MD can also be used to introduce soft geometric constraints in which the controlled geometric parameter is not strictly constant. Instead, it oscillates in a narrow interval of values.
In VASP, the bias potentials are supported in both the NVT and NpT MD simulations, regardless of the particular thermostat and/or barostat setting. Different types of potentials can be freely combined. The values of all collective variables defined in the ICONST file for each MD step are listed in the REPORT file. Check the lines after the string Metadynamics.
Theoretical background
The probability density for a geometric parameter ξ of the system driven by a Hamiltonian:
with T(p), and V(q) being kinetic, and potential energies, respectively, can be written as:
The term stands for a thermal average of quantity X evaluated for the system driven by the Hamiltonian H.
If the system is modified by adding a bias potential acting on one or multiple selected internal coordinates of the system ξ=ξ(q), the Hamiltonian takes a form:
and the probability density of ξ in the biased ensemble is:
It can be shown that the biased and unbiased averages are related via a simple formula:
More generally, an observable :
can be expressed in terms of thermal averages within the biased ensemble:
Simulation methods, e.g., umbrella sampling[1], use a bias potential to enhance sampling of ξ in regions with low P(ξi), e.g., transition regions of chemical reactions. The correct distributions are recovered afterward using the equation for above. Ref.[2] provides a more detailed description of the method.
Supported types of bias potentials
Presently, the following types of bias potential are supported:
Harmonic potentials
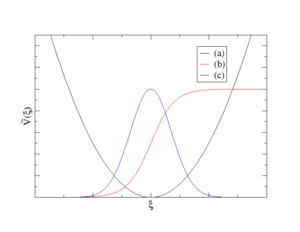
- A sum of Harmonic potentials (see curve (a) on the plot above)
- where the sum runs over all () coordinates the potential acts upon. The coordinates are defined in the ICONST file by setting the
statusto 8. The parameters of the potential are the force constant (SPRING_K) and the minimum or the potential SPRING_R0. These must be set in the INCAR file. Optionally, it is possible to change the value of every MD step at a constant rate defined via the INCAR tag SPRING_V0.
Mind: The number of items defined via SPRING_K, SPRING_R0, and SPRING_V0 must be equal to . Otherwise, the calculation terminates with an error message.
- This form of bias potential is employed in several simulation protocols, such as the umbrella sampling[1], umbrella integration, or steered MD, and is useful also in cases where the values need to be restrained.
Step function
- A sum of Fermi-like step functions (see curve (b) on the plot above)
- where the sum runs over all () coordinates the potential acts upon. The coordinates are defined in the ICONST file by setting the
statusto 4. The parameters of the potential are the height of the step ( set by FBIAS_A), the slope around the point ( set by FBIAS_D), and the position of the step ( set by FBIAS_R0). These must be set in the INCAR file.
Mind: The number of items defined via FBIAS_A, FBIAS_D, and FBIAS_R0 must be equal to . Otherwise, the calculation terminates with an error message.
- This form of potential is suitable especially for imposing restrictions on the upper (or lower) limit of the value of .
Gaussian potential
- A sum of Gauss functions (see curve (b) on the plot above)
- where is the number of Gaussian functions and is the number of coordinates the potential acts upon. The coordinates are defined in the ICONST file by setting the
statusto 5. The parameters of the potentials, , , and are defined in the PENALTYPOT file.
- This type of bias potential is primarily intended for use in metadynamics, but since Gaussians can be used as basis functions for more general shapes, they can also be used to prepare various atypically-shaped bias potentials.
Examples of usage
Let us consider the nucleophile substitution reaction of CHCl with Cl. The reactant is a weak van-der-Waals complex. The corresponding POSCAR file reads
vdW complex CH3Cl...Cl 1.00000000000000 12.0000000000000000 0.0000000000000000 0.0000000000000000 0.0000000000000000 12.0000000000000000 0.0000000000000000 0.0000000000000000 0.0000000000000000 12.0000000000000000 C H Cl 1 3 2 cart 5.91331371 7.11364924 5.78037960 5.81982231 8.15982106 5.46969017 4.92222130 6.65954232 5.88978969 6.47810398 7.03808479 6.71586385 4.32824726 8.75151396 7.80743202 6.84157897 6.18713289 4.46842049
Due to the weak interactions between CHCl and Cl, the complex can collapse at high temperatures. This can be avoided by setting an upper bound for the length of the non-bonding Cl...C interactions. This can be conveniently achieved by using a Fermi-like step-shaped bias potential. To this end, we need to define the Cl...C distance, i.e., the distance between the atoms 1 and 5, as a coordinate with status 4 in the ICONST file:
R 1 5 4
Next, we need to specify the bias potential parameters FBIAS_A, FBIAS_D, and FBIAS_R0 in the INCAR file. Since the bias potential acts only on one internal coordinate (), we need to provide only one number for each of the tags. The setting
FBIAS_A = 1 FBIAS_D = 50 FBIAS_R0 = 3.5
ensures that repulsive bias forces steeply increase when the C...Cl distance is increased beyond about . This causes a shortening of the distance in the next MD step. Notice that the bias force is essentially negligible for distances below . A careful adjustment of FBIAS_A and FBIAS_D is needed to ensure that (i) the bias force is large enough to effectively limit the value of , and (ii) the interval of values for which the bias forces are significant is broad enough to avoid overcoming via random fluctuations. A suitable setting can be found by noting that the maximal bias force of is exerted on the system at the point . This can be seen by inspecting the analytical expression for the potential.
How to
Biased MD run using the Andersen thermostat and a Gaussian bias potential
- For a biased MD run with Andersen thermostat, one has to:
- Set the standard MD-related tags: IBRION=0, TEBEG, POTIM, and NSW
- Set MDALGO=1 (MDALGO=11 in VASP 5.x), and choose an appropriate setting for ANDERSEN_PROB
- To avoid updating of the bias potential, set HILLS_BIN=NSW
- Define collective variables in the ICONST file, and set the STATUS parameter for the collective variables to 5
- Define the bias potential in the PENALTYPOT file
Biased MD run using the Nose-Hoover thermostat and a Gaussian bias potential
- For a biased MD run with Nose-Hoover thermostat, one has to:
- Set the standard MD-related tags: IBRION=0, TEBEG, POTIM, and NSW
- Set MDALGO=2 (MDALGO=21 in VASP 5.x), and choose an appropriate setting for SMASS
- To avoid updating the bias potential, set HILLS_BIN=NSW
- Define collective variables in the ICONST file, and set the STATUS parameter for the collective variables to 5
- Define the bias potential in the PENALTYPOT file
The values of all collective variables for each MD step are listed in the REPORT file. Check the lines after the string Metadynamics.
References
This category currently contains no pages or media.
